The health weather app
The app
Forecasting weather is one of the most widespread products in any form of media. Having a broad idea of how the weather is going to be throughout the day can help us to better plan our activities. From changing our outfits to even preventing catastrophes, weather forecasting is a powerful tool that changes our lives for the better.
Like weather, some contagious diseases also follow recurrent seasonal patterns. Often, such patterns can appear random and unpredictable. Still, the use of state-of-the-art technology can help us to better understand the nature of such variations.
The Health Weather app is an Android app that leverages the power of generative models to forecast the prevalence of COVID-19. It also provides a genome forecast, enabling its user to stay up to date on the most likely genomic variant. Every single component inside the neural engine is shipped inside the app. Allowing the user to access the forecast even without an internet or mobile connection.
This beta release aims to raise funds to continue the development of the app. The long-term goal is to add other diseases like influenza, dengue, hepatitis, and many other viral diseases. As well as to make the models smaller and more accurate.
The UI
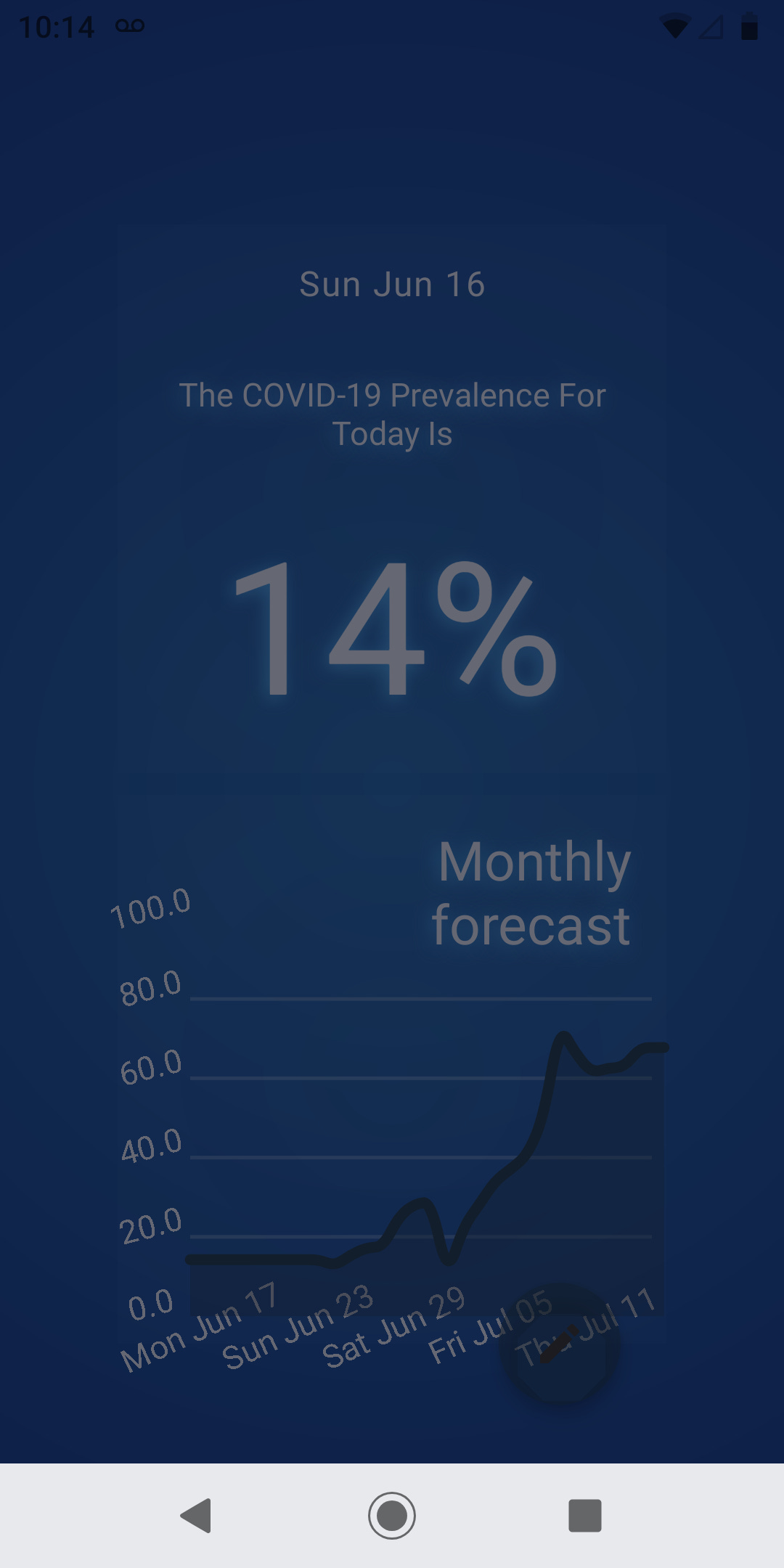
The interface contains three main elements, a couple of panels and a button. The first panel displays the daily forecast. This percentage value represents the forecasted prevalence of COVID-19 at the present day. The second panel displays a graph with the monthly forecast value. This second panel shows the forecasted prevalence for the following 30 days.
The button on the bottom allows the user to copy the forecasted genome into the clipboard. Daily genomes can be saved using a text editor or a note-taking app.
How the forecast is made?
The forecast is divided into two estimators, a density estimator and a machine learning estimator. The first one approximates the prevalence of SARS-Cov2 by the density of isolated genomes through the PHEIC phase of the pandemic. The second one relies on the daily genome prediction. And the correlation between the amount of adenine inside the SARS-Cov2 genome and the prevalence of COVID-19. You can find an in-depth analysis of the SARS-Cov2 genome in the following link.
The genome is predicted using a local deep 3D convolutional network and the amount of adenine is measured from that prediction. Both estimations are added together and displayed in the graph and main score.
From the two estimators, the genomic one is updated daily while the density one is a fixed dataset. Thus the density estimation can be interpreted as the baseline forecast. While the daily update is a correction that increases or lowers that baseline.
Testing the forecasted genome
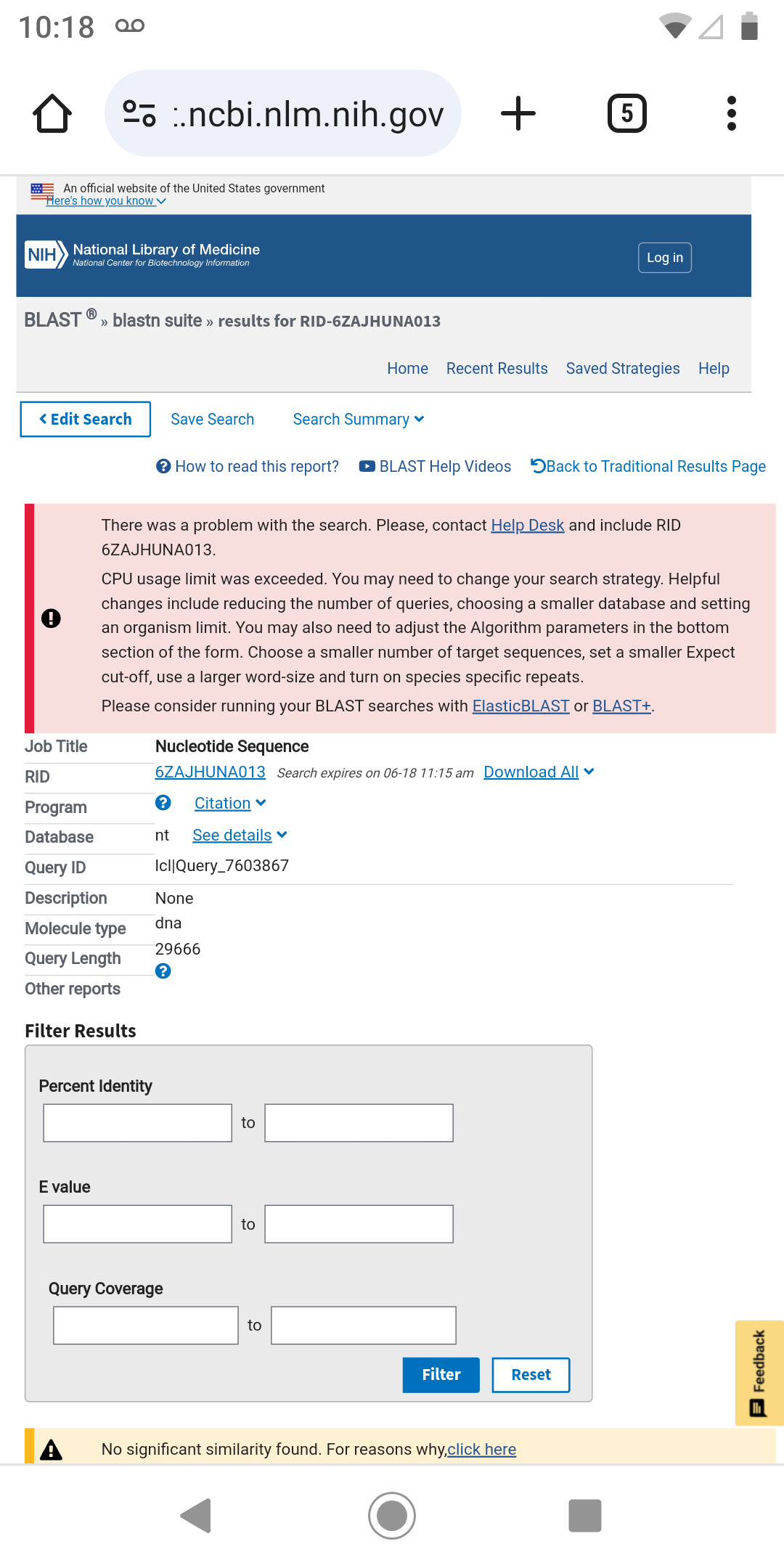
You can copy to the clipboard the forecasted genome to the clipboard by using the pen icon on the bottom left. To test the identity of the forecasted genome you can use the BLAST service from NCBI. But to ensure the identification of the genome you should provide only a fraction of the forecasted genome. Using the complete one will most likely result in a CPU usage error.
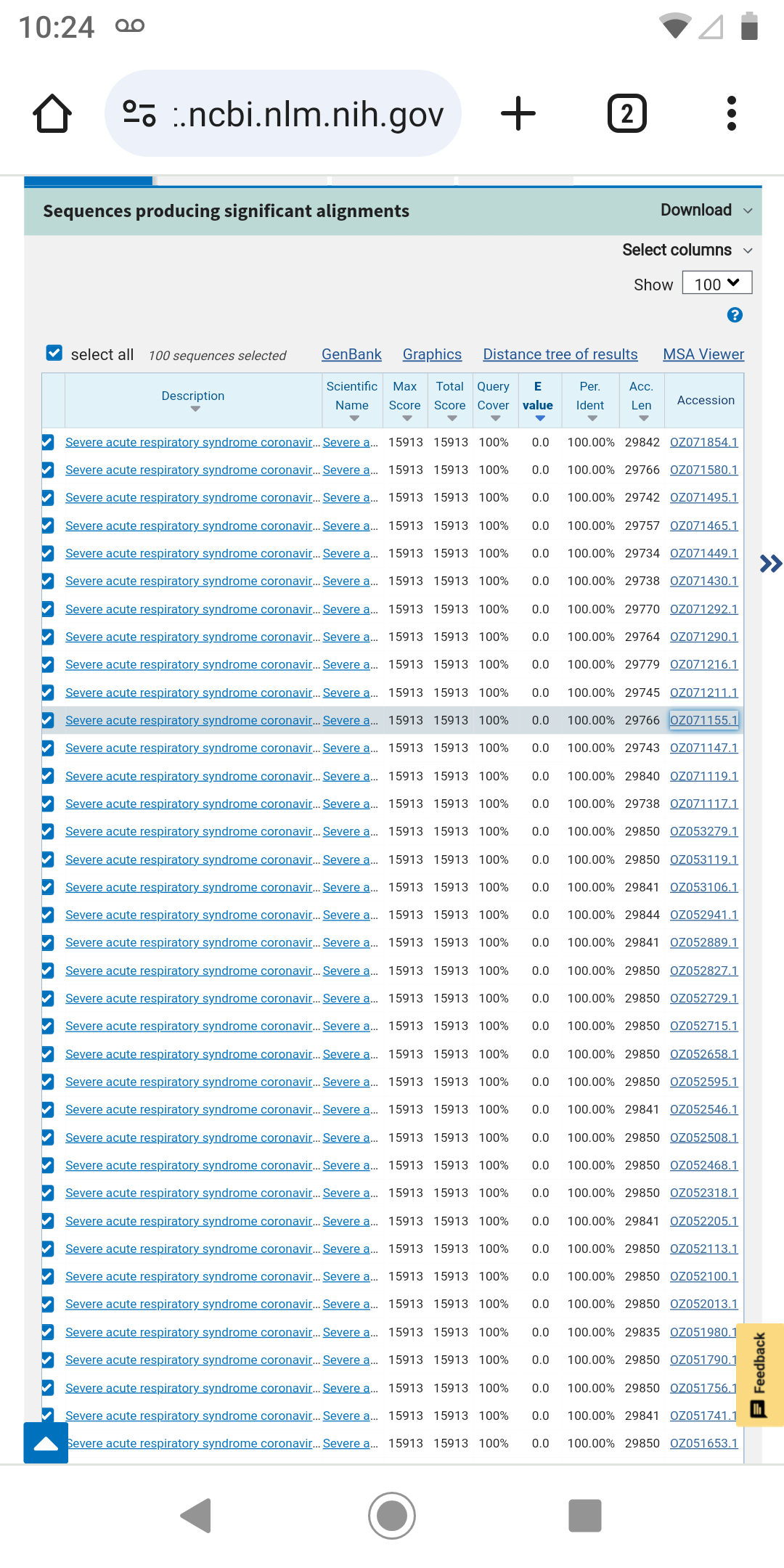
Using a fraction of the genome the BLAST service can find a series of SARS Cov2 genomes like the one forecasted by the app.
The forecasted genome will be the same for 24 hours after that period the forecast is updated.
You can find the beta version of the app on Gumroad or buymeacoffe.