Viruses as dynamical systems
A hallmark characteristic of viruses is their ability to change. Changes in the genetic information of viruses are known as mutations. Some mutations will have no impact on the viral genes leading to an exact copy of the expressed genes. Other mutations will change a single or multiple proteins expressed by the virus. Leading to either an equally-adapted virus, a better-adapted one, or a defective virus incapable of replicating itself.
Regardless of the kind of mutation, changes in the viral genome will change the frequency of each of the nucleotides that make up the genome. The size of those changes will depend on the mutation rate of the virus. While our ability to detect those changes will depend on how frequently the virus is being sampled. Thus low mutation rate and scarcely sampled virus could appear as stable. While a scarcely sampled virus with a high mutation rate will appear as unpredictable.
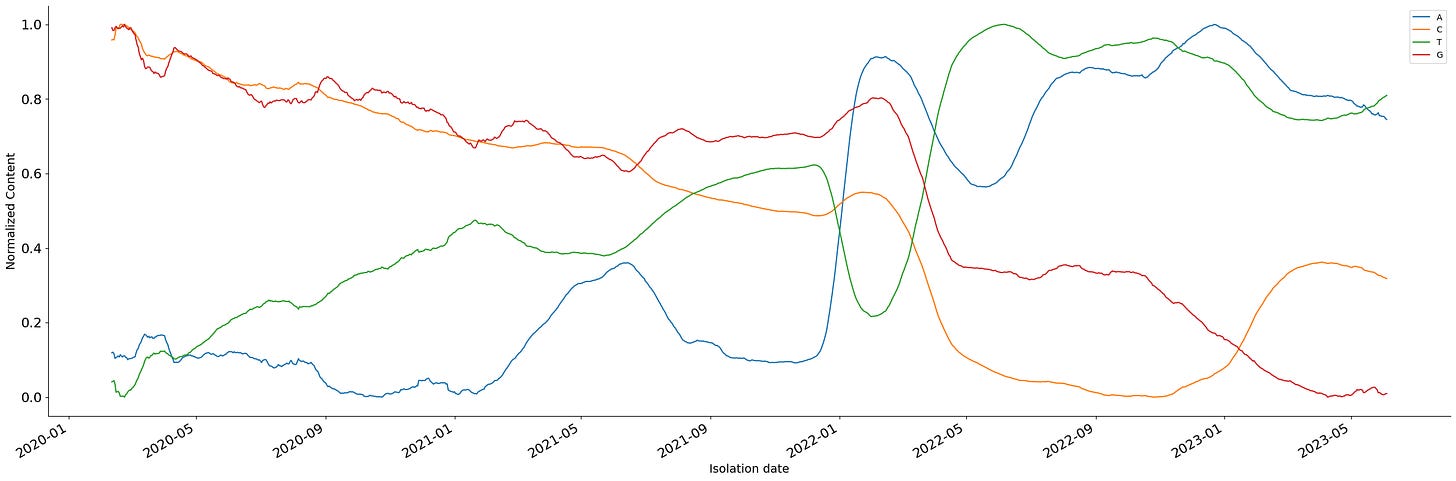
But, even understanding the behavior of frequently sampled viruses could be a challenging task. Let's take SARS-Cov2, the most sampled virus in human history at the moment. If we plot the relative frequency of each of the nucleotides by time we can see a series of oscillations through time.
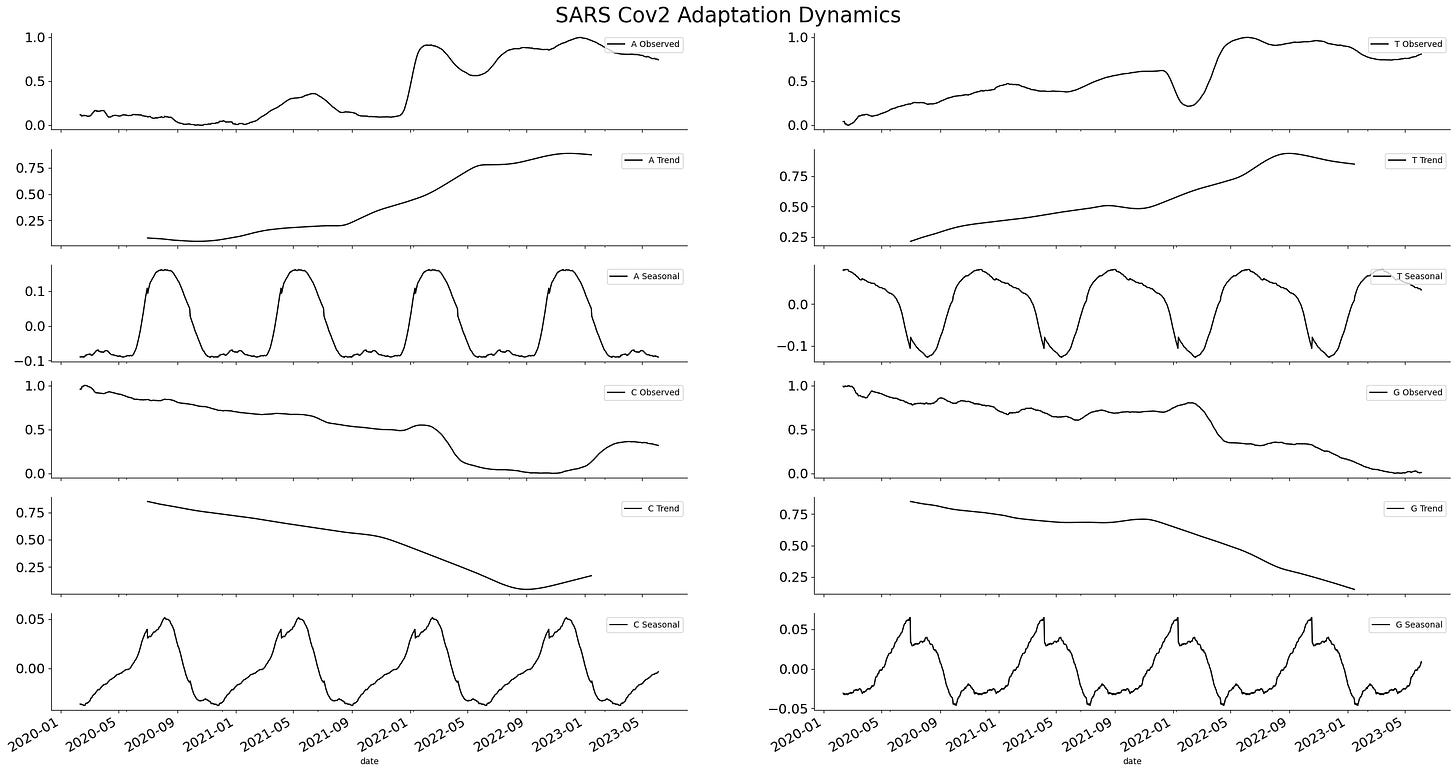
This simple time series can be decomposed into a combination of two elements. The first one describes the long-term trajectory of the time series and is known as the trend component. The second describes the oscillation inside the time series known as the seasonal component.
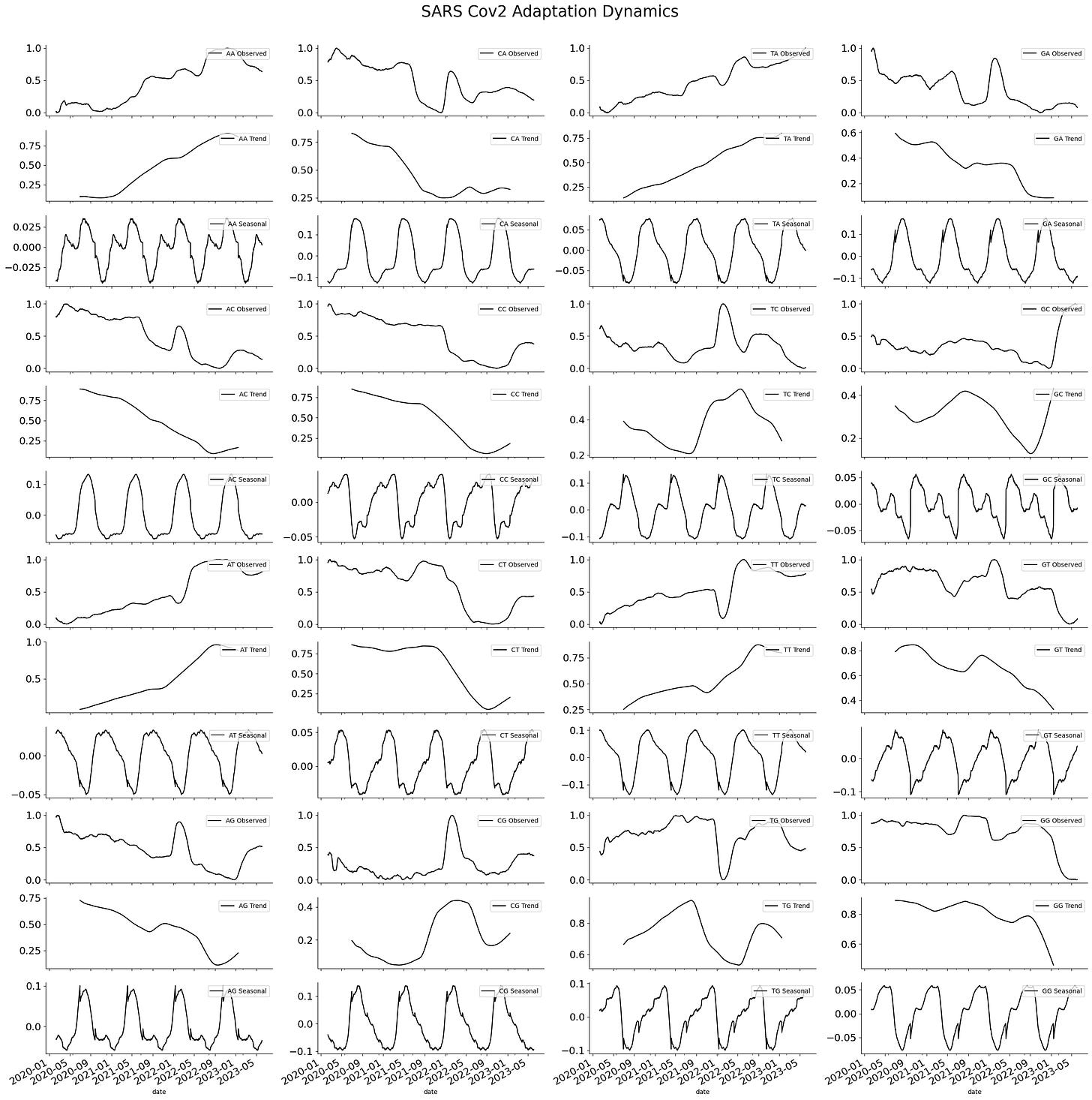
This analysis shows that changes in the frequency of nucleotides follow a seasonal pattern. Expanding the analysis to the frequency of dinucleotides also shows a similar pattern. A series of seasonal elements are encoded inside the viral genome.
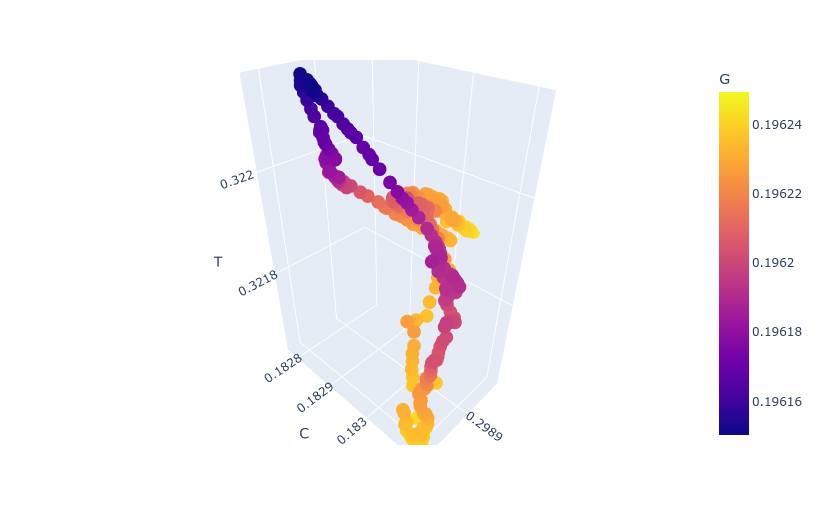
However, analyzing both components can prevent us from finding meaningful patterns inside the data. If we take only the seasonal component with a duration of roughly one year we can try to reconstruct the path followed by the virus. Plotting the frequency of of nucleotides on the x,y, and z axes and encoding the remaining one as color we get a cyclical path.
This path represents the mean trajectory followed by the virus and most likely the mean representation of the attractor generated by analyzing genomes as a time series. This also hints that the frequency of nucleotides inside SARS-Cov2 can be described as a system of differential equations. Finding such a system will most likely help us to better understand the long and short-term dynamics inside the virus.
Even though the previous analysis deviates from the traditional one, I will argue that both analyses are not incompatible. A specific virus could be represented mathematically as a single attractor. Extreme regions inside the attractor could represent different branches inside the phylogenetic tree. Similar regions will share phenotypic characteristics as well as recurrent genomic sequences. Variation inside the genome could be a consequence of the virus orbiting the attractor. In theory, every single virus will be able to generate any other possible kind of variant.
The attractor/dynamical system representation of a virus could also provide a continuous description of the ability of a virus to replicate itself given a set of conditions. Yielding a quantitative representation of its fitness landscape. The current mean attractor representation for SARS Cov2 already encodes information about its ability to infect other hosts through time. Time values can be transformed into different environmental variables defining the attractor as a parametric curve. This new representation could point to specific mutations or adaptations specific to certain environmental conditions.
Add an extra layer of protection and prevent COVID-19 infections, try the health weather app beta for free. Available on Gumroad.